Our gene expression is regulated by RNA methylation. Same process also affects various cellular processes like differentiation and development. Lately, University of Chicago researchers, led by Chuan He, have uncovered new insights into how genes work by solving a long-standing mystery related to RNA methylation.
Researchers in Chuan He’s lab have been studying RNA methylation and its potential impact on human health for over a decade.
Until, 20th century we believed that DNA (deoxyribonucleic acid) contains the genetic information that serves as the blueprint for the development and function of cells. And therefore, everything is “pre-determined”.
This, however, is not the real picture. Our bodies are more susceptible to our experiences and environment. And can turn some genes on and off as per requirement. For instance, environment guides the growth of a plant – it may grow shorter relatively in times of drought. When we expose our skin to sun light for long, it tends to generate more melanin for protection.
RNA methylation plays an important role in developing such environmental responses.
RNA methylation
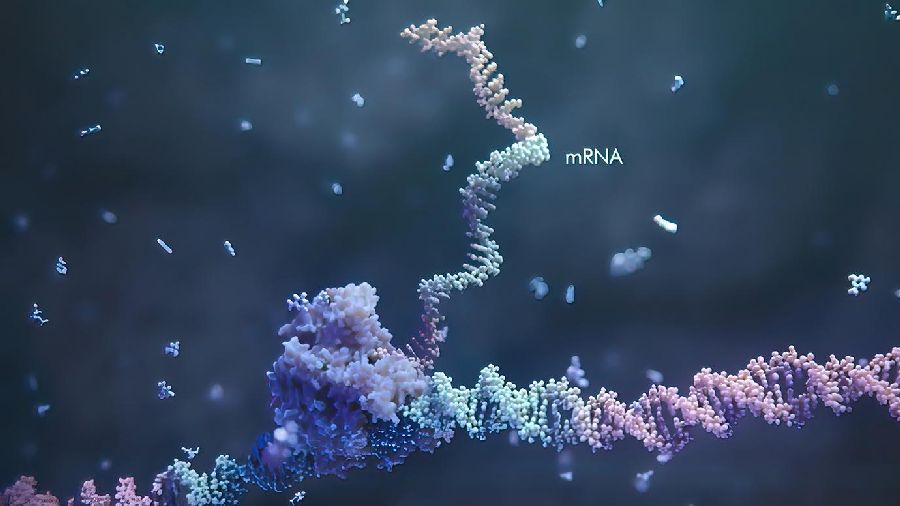
RNA (ribonucleic acid), generally, copies DNA and carries instructions to make different proteins in the cell. At times, while carrying genetic information from DNA to the ribosomes, RNA changes those instructions.
One of the ways to alter the gene expression is by adding a methyl group to the RNA molecule. This modification changes the instruction that is suppose to carried out. Thus, the entire process influences the expression of the gene it encodes.
Chuan He highlighted this significance of RNA methylation in controlling cell differentiation, where specific cells develop into different cell types like skin, muscle, or eyes.
RNA methylation is to select site where not to methylate
However, the underlying mechanisms of RNA methylation is still not known. Not all but only “certain sites” on the RNA molecule can be methylated and the reason remained unclear.
He’s research group discovered that cells do not selectively choose specific sites to methylate during RNA modification rather, they pick where not to methylate. And the joints of messenger RNA play a vital role in this process.
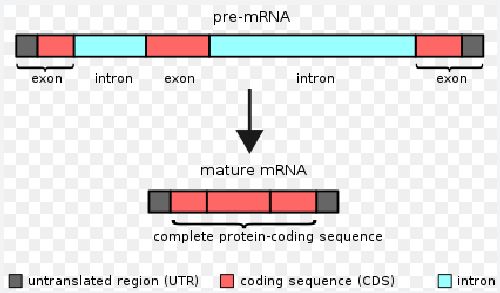
RNA Splicing
After RNA copies DNA, the RNA undergoes a process called splicing in which some parts of the messenger RNA molecule are removed, and the remaining parts are joined together.
The editing occurs because the discarded regions do not code for translated regions. Non coding regions called introns, and the coding regions called exons. After removing the introns, the exons are joined together to form a continuous RNA molecule.
RNA splicing produces mature mRNA molecules that are used as templates for protein synthesis.
Exon junction complex
The precise mechanisms and functions of the “exon junction complex” are an active area of research in molecular biology including Chuan’s lab.
The team discovered that the exon junction complex plays a role in determining whether specific sites on messenger RNA molecules can be methylated or not. For instance, bulky molecules at the ends of short RNA pieces inhibit methylation while longer pieces with more space between them expose specific sites to methylation.
Takeaway
Researchers concluded that the length and spacing of the RNA molecule could affect gene expression. Therefore, the discovery may have implications for the development of artificial genes used in gene therapies and basic research.
This is an interesting study surfacing the complex processes that govern gene expression. With this insight, scientists will surely be able to bring about more effective gene therapies.